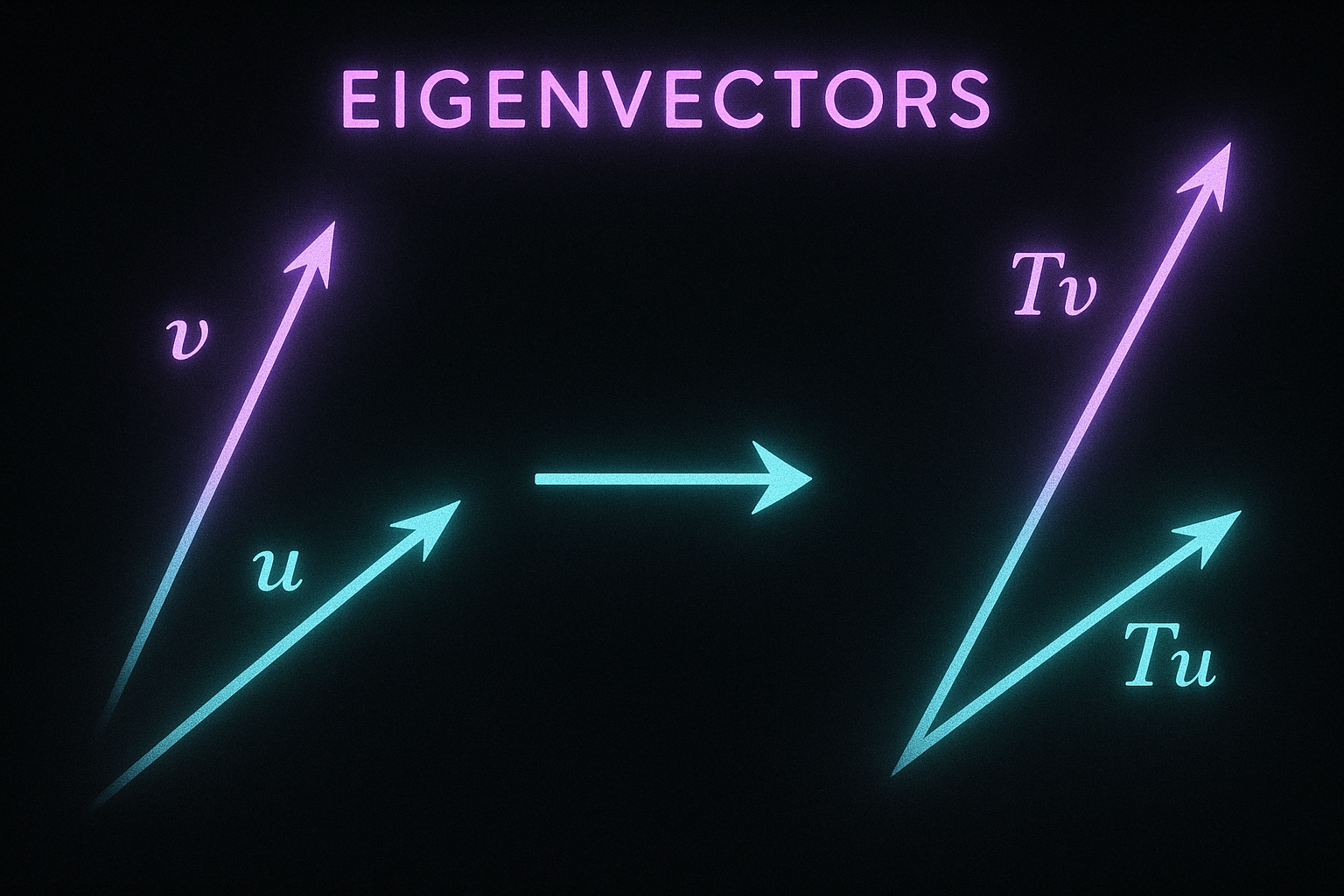
A matrix transforms vectors. Eigenvectors are special directions that only get scaled, not rotated. Eigenvalues tell you the scale factor.
Sounds abstract but used everywhere - PCA, spectral clustering, stability analysis, PageRank.
Definition
For matrix A, eigenvector v and eigenvalue λ satisfy:
$$Av = \lambda v$$
Apply matrix → just multiplies by scalar. Direction unchanged.
Interactive demo: Eigenvalue Animation
Finding eigenvalues
Rewrite: Av = λv → (A - λI)v = 0
For non-trivial v, matrix (A - λI) must be singular:
$$\det(A - \lambda I) = 0$$
This gives the characteristic polynomial. Roots are eigenvalues.
For 2×2 matrix:
A = [a b]
[c d]
det(A - λI) = (a-λ)(d-λ) - bc = λ² - (a+d)λ + (ad-bc) = 0
Computing in practice
Don’t solve characteristic polynomial for large matrices. Use numerical methods.
import numpy as np
A = np.array([[4, 2], [1, 3]])
# Eigenvalues and eigenvectors
eigenvalues, eigenvectors = np.linalg.eig(A)
print(eigenvalues) # [5. 2.]
print(eigenvectors) # columns are eigenvectors
Properties
Real symmetric matrices:
- All eigenvalues are real
- Eigenvectors are orthogonal
- Can be diagonalized
Trace = sum of eigenvalues: $$\text{tr}(A) = \sum \lambda_i$$
Determinant = product of eigenvalues: $$\det(A) = \prod \lambda_i$$
PCA connection
Principal Component Analysis uses eigenvalues.
- Compute covariance matrix of data
- Find eigenvectors (principal components)
- Eigenvalues tell variance explained
# PCA from scratch
X_centered = X - X.mean(axis=0)
cov = X_centered.T @ X_centered / len(X)
eigenvalues, eigenvectors = np.linalg.eigh(cov)
# Sort by eigenvalue (descending)
idx = np.argsort(eigenvalues)[::-1]
principal_components = eigenvectors[:, idx]
Or just use sklearn:
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
X_reduced = pca.fit_transform(X)
Spectral decomposition
If A is symmetric, it can be written as:
$$A = Q\Lambda Q^T$$
Where:
- Q has eigenvectors as columns
- Λ is diagonal matrix of eigenvalues
This is powerful for matrix analysis.
Power iteration
Simple algorithm to find largest eigenvalue:
def power_iteration(A, num_iters=100):
v = np.random.randn(A.shape[0])
v = v / np.linalg.norm(v)
for _ in range(num_iters):
v = A @ v
v = v / np.linalg.norm(v)
eigenvalue = v @ A @ v
return eigenvalue, v
Converges to dominant eigenvector. Used in PageRank!
Eigenvalues and stability
For differential equations dx/dt = Ax:
- All eigenvalues negative → stable (decays)
- Any eigenvalue positive → unstable (grows)
- Complex eigenvalues → oscillation
In neural networks, eigenvalues of weight matrices relate to gradient flow.
Condition number
Ratio of largest to smallest eigenvalue (for symmetric positive definite):
$$\kappa(A) = \frac{\lambda_{max}}{\lambda_{min}}$$
Large condition number = ill-conditioned = numerical problems.
cond = np.linalg.cond(A)
# If very large, matrix is nearly singular
Eigenvalues clicked? Show your support by starring ML Animations on GitHub and sharing on LinkedIn!