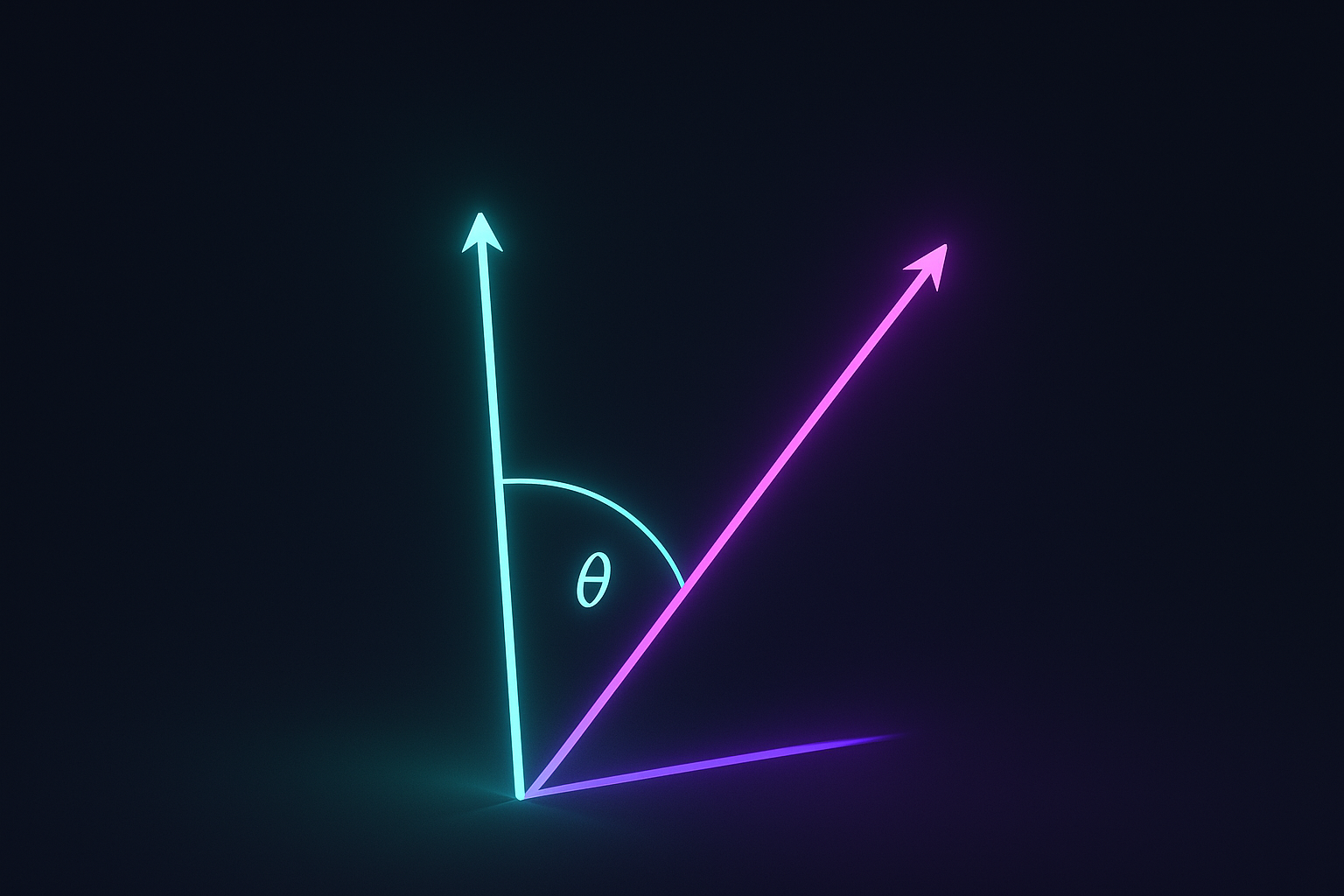
Two embeddings point in similar direction? They’re similar. Cosine similarity ignores magnitude, only cares about angle.
Definition
$$\cos(\theta) = \frac{\mathbf{a} \cdot \mathbf{b}}{|\mathbf{a}| \cdot |\mathbf{b}|}$$
Range: -1 (opposite) to 1 (same direction). 0 means perpendicular (orthogonal).
Interactive demo: Cosine Similarity Animation
Computing it
import numpy as np
def cosine_similarity(a, b):
return np.dot(a, b) / (np.linalg.norm(a) * np.linalg.norm(b))
# Or with scipy
from scipy.spatial.distance import cosine
similarity = 1 - cosine(a, b) # cosine returns distance
# Or sklearn for batches
from sklearn.metrics.pairwise import cosine_similarity
sims = cosine_similarity(X, Y) # pairwise similarities
Why not Euclidean?
Consider embeddings:
- “king”: [0.5, 0.5]
- “queen”: [0.4, 0.4]
- “peasant”: [0.1, 0.1]
Euclidean says peasant is closer to queen than king is! Cosine says king ≈ queen ≈ peasant (all same direction).
For normalized embeddings (unit vectors), Euclidean and cosine give same ranking.
When to use
Cosine:
- Text/word embeddings
- When magnitude is noise
- High-dimensional sparse data
Euclidean:
- Physical distance
- When magnitude matters
- Low-dimensional dense data
In information retrieval
TF-IDF vectors are high-dimensional and sparse. Cosine similarity is standard:
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
vectorizer = TfidfVectorizer()
tfidf = vectorizer.fit_transform(documents)
# Find most similar to query
query_vec = vectorizer.transform([query])
similarities = cosine_similarity(query_vec, tfidf)
most_similar = np.argsort(similarities[0])[::-1]
For embeddings search
RAG, semantic search, all use cosine similarity:
# Query embedding
query_emb = model.encode("what is machine learning")
# Find similar in database
similarities = cosine_similarity([query_emb], document_embeddings)[0]
top_k = np.argsort(similarities)[-5:][::-1]
Cosine distance
Distance = 1 - similarity
from scipy.spatial.distance import cosine
distance = cosine(a, b) # 0 = same, 2 = opposite
Sometimes you need distance (clustering, kNN) not similarity.
Batched computation
For efficiency, compute many similarities at once:
# Normalize once
X_norm = X / np.linalg.norm(X, axis=1, keepdims=True)
Y_norm = Y / np.linalg.norm(Y, axis=1, keepdims=True)
# All pairwise similarities
similarities = X_norm @ Y_norm.T
Matrix multiply normalized vectors = cosine similarity matrix.
Soft cosine similarity
Account for word similarity in bag-of-words:
$$\text{soft_cos}(a, b) = \frac{a^T S b}{\sqrt{a^T S a} \sqrt{b^T S b}}$$
Where S is word similarity matrix. “Happy” and “joyful” get credit for being similar.
In neural networks
Cosine similarity as loss component:
import torch.nn.functional as F
# Cosine similarity
sim = F.cosine_similarity(a, b)
# Cosine embedding loss (for similar/dissimilar pairs)
loss = F.cosine_embedding_loss(a, b, target) # target: 1 or -1
Contrastive learning
CLIP, SimCLR use cosine similarity:
# Simplified contrastive loss
def contrastive_loss(anchor, positive, negatives, temperature=0.1):
pos_sim = F.cosine_similarity(anchor, positive) / temperature
neg_sims = F.cosine_similarity(anchor.unsqueeze(1), negatives) / temperature
logits = torch.cat([pos_sim.unsqueeze(1), neg_sims], dim=1)
labels = torch.zeros(len(anchor), dtype=torch.long)
return F.cross_entropy(logits, labels)
Like what you learned? Help others discover these concepts by starring ML Animations on GitHub and sharing on LinkedIn or Twitter!